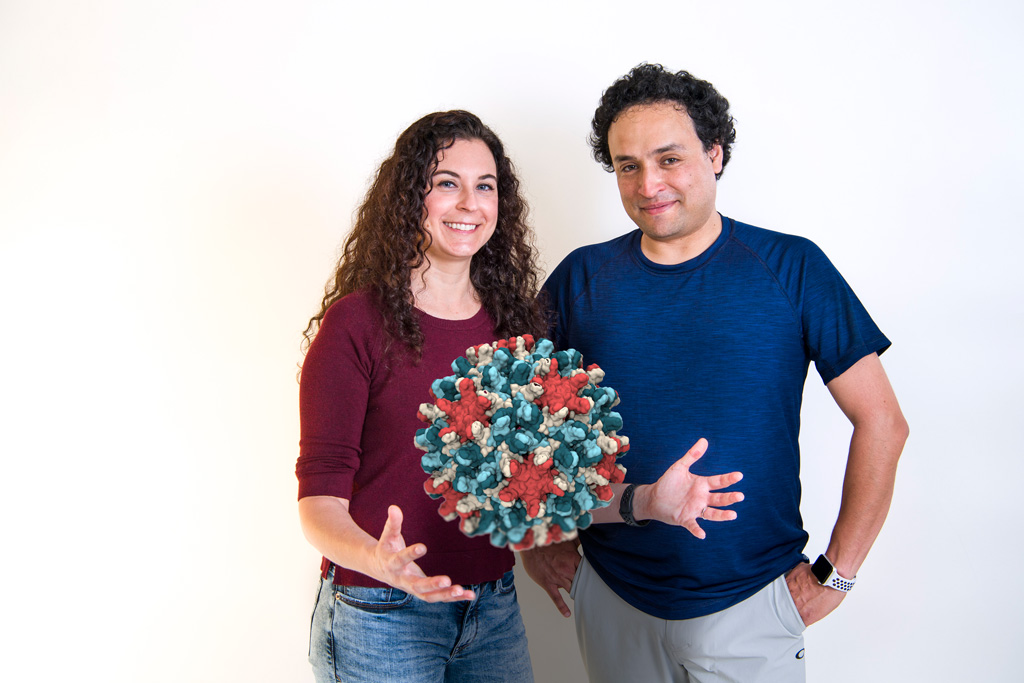



A view of a virus
Photo by Kathy F. Atkinson | Photo illustration by Jeffrey Chase May 31, 2018
Research reveals new insights into hepatitis B
Researchers at the University of Delaware, working with colleagues at Indiana University, have gained new insights into the virus that causes hepatitis B — a life-threatening and incurable infection that afflicts more than 250 million people worldwide.
The discovery, which was published April 27 in the journal eLife, reveals previously unknown details about the capsid, or protein shell, that encloses the virus’ genetic blueprint.
Scientists believe that the capsid, which drives the delivery of that blueprint to infect a host cell, is a key target in developing drugs to treat hepatitis B.
“With hepatitis B, the structure of the capsid has been known for years, but we wanted to study its motion and its influence on its surroundings,” said Jodi A. Hadden, an independent postdoctoral researcher in UD’s Department of Chemistry and Biochemistry and the lead author of the new paper.
Hadden and the research team used supercomputing resources to perform what are known as all-atom molecular dynamics simulations.
Molecular dynamics simulations allow researchers to study the way molecules move in order to learn how they carry out their functions in nature. Computer simulations are the only method that can reveal the motion of molecular systems down to the atomic level and are sometimes referred to as the “computational microscope.”
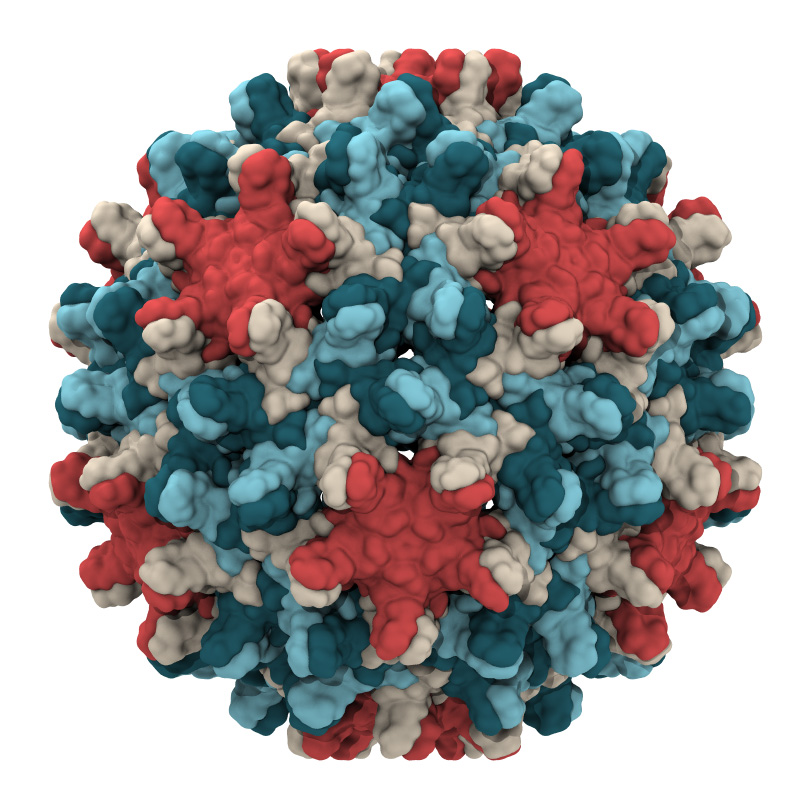
In the case of the simulations of the hepatitis B virus, the researchers found that the capsid is not rigid as previously thought, but is highly flexible. They also learned that it can distort into an asymmetric shape, which might allow it to squeeze through an opening into the nucleus of a cell the virus is infecting.
“We think that the capsid might need that ability to distort in order to correctly package its genetic blueprint and get it into the nucleus to generate new copies of the virus during the infection process,” Hadden said.
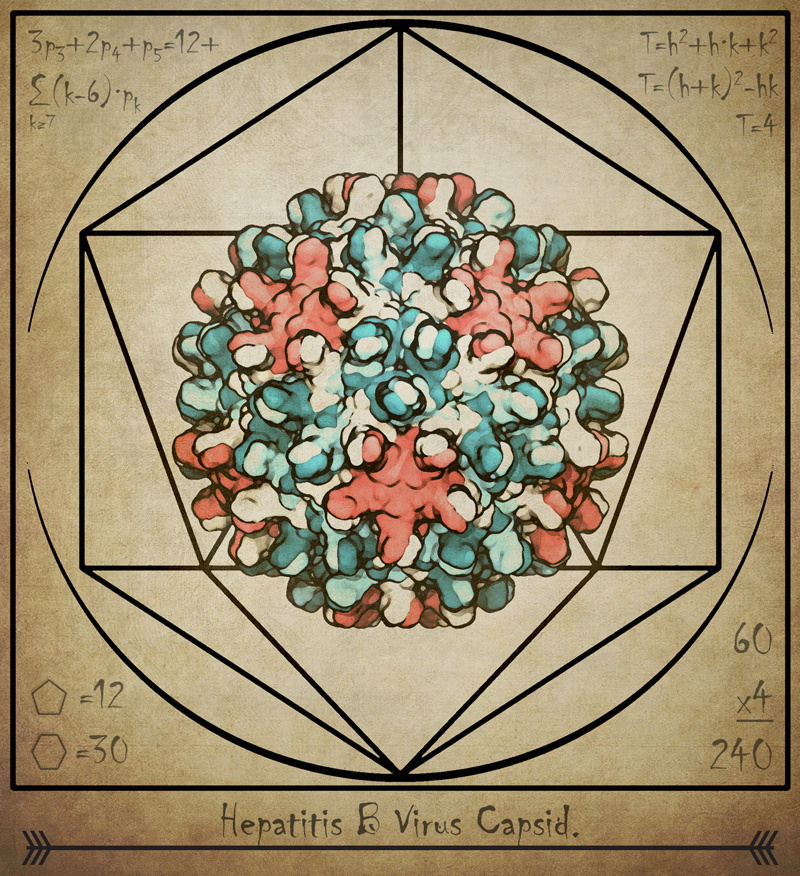
Previous research has used experimental microscopes to study the capsid, which is made up of 240 proteins, but that work hasn’t yielded high-resolution images of the complex structure, said Juan R. Perilla, assistant professor of chemistry and biochemistry and a co-author of the new paper.
“It seems clear that the flexibility of the [hepatitis B] capsid is a limiting factor” in the effectiveness of microscopy, he said.
By contrast, the simulations have been able to reveal a more complete picture of the capsid and how it moves, distorts and interacts with its environment, Hadden said. Each simulation involves six million atoms.
“We have all the details down to the atomic level,” she said. “You need that to develop a complete understanding of the molecule and to study drug interactions.”
The researchers also found that small triangular openings, or pores, in the capsid surface are likely the location where its protein “tails” poke through, sending a signal that is essential to the infection process.
“We know that the capsid tails have to be exposed to the surface at some time for the capsid to travel to the cell nucleus,” Hadden said. “It’s like hailing a taxi.”
All the findings have the potential to lead to drug treatments, she said. For example, if the capsid could be made rigid and unable to distort or if a way could be found to block the triangular pores in its surface, the infection process might be halted.
There’s an effective vaccine to prevent hepatitis B, but no cure once a person is infected. The virus causes severe liver disease, which can lead to potentially fatal conditions such as cirrhosis and liver cancer.
More about the research
The paper is available online at eLife, a peer-reviewed, open-access scientific journal for the biomedical and life sciences.
Co-authors, with Hadden and Perilla, are Christopher John Schlicksup, Balasubramanian Venkatakrishnan and Adam Zlotnick, all of Indiana University, and the late Klaus Schulten of the University of Illinois at Urbana-Champaign (UIUC), who pioneered the application of all-atom molecular dynamic simulations to study complete virus capsids.
The research was supported by UD, through a postdoctoral fellowship to Hadden, and the National Institutes of Health, through a Center of Biomedical Research Excellence Grant to Perilla and a Biomedical Technology Research Resource to Schulten. The computer simulations were made possible by the Blue Waters project, a joint effort of UIUC and its National Center for Supercomputing Applications.
Contact Us
Have a UDaily story idea?
Contact us at ocm@udel.edu
Members of the press
Contact us at mediarelations@udel.edu or visit the Media Relations website

