


The genetics of cancer
Photo by Kathy F. Atkinson | Photo illustration by Jeffrey C. Chase September 12, 2019
UD molecular biologist and colleagues reveal new insights into tumor progression
University of Delaware molecular biologist Mona Batish and collaborators at Harvard Medical School and University of California, Los Angeles, have identified a new circular ribonucleic acid (RNA) that increases tumor activity in soft tissue and connective tissue tumors.
Finding this new genetic unit has the potential to advance understanding of the genetics of cancer and how cancer is identified and treated.
The researchers recently reported their findings in a new paper in Cell Research, a Nature journal. Batish was a co-author on the team that included Jlenia Guarnerio, the paper’s lead author and assistant professor of biomedical sciences at UCLA and at Cedars-Sinai Medical Center; Pier Paolo Pandolfi, the Aresty Endowed Chair of Medicine and professor of medicine and pathology at Harvard Medical School; and colleagues from Harvard Medical School’s Beth Israel Deaconess Medical Center, Rutgers University and Aalborg University Hospital in Denmark.
A word about circular RNA
RNA is a single-stranded molecule that is made by the DNA — the code of life — in our bodies. Messenger RNA (mRNA) acts as a courier, transporting instructions from the DNA code to protein-making machines and thus dictating the composition of proteins in a cell. Apart from mRNA, there are many other types of RNA, which do not carry code for proteins but perform other important functions in cells. Collectively, these are known as non-coding RNAs.
A new class of non-coding RNA, called circular RNA, was discovered in the 1970s. Circular RNA (circRNA) was initially thought to be a virus because most RNA molecules are linear, meaning that their genetic sequence always moves in a forward direction. By contrast, circRNA is circular, even though it shares the same genetic sequence as linear RNA.
“Under certain conditions, RNA processing systems can get tricked into thinking they are supposed to join the ends,” said Batish, an assistant professor of medical and molecular sciences in UD’s College of Health Sciences. “When this error occurs, it creates a backwards loop in the RNA’s genetic sequence and then keeps on going— kind of like when you get a kink in the middle of a necklace.” This loop separates off and persists as a circular RNA inside the cell.
For a long time, researchers thought this error, a process known as back splicing, didn’t mean anything. But when genome sequencing came about in the 1990s, scientists started finding circular RNA in brain tissue and other tissues. By 2014, they realized that circular RNA was important and, today, there is a whole field looking at circular RNA as a biomarker for disease, particularly cancers.
According to Batish, the role of circRNA in tumor progression has been understudied.
In the paper, the researchers describe a new circRNA generated by a gene called Zbtb7a found in soft tissue tumors, such as mesenchymal tumors. In its linear form, this RNA makes a tumor-suppressing protein that stops the growth of cancer, according to previous research out of Pandolfi’s Harvard research lab. However, once the same RNA makes a circRNA (that is, gets a “kink”), the circular RNA works independently to make the tumor more active, effectively silencing the tumor-suppressing protein.
According to Batish, this is the first time that this type of antagonistic, tumor-promoting role of circRNA has been shown in connection with linear RNA with the same genetic sequence.
Theoretically, both RNA strands should perform the same function because they originate from the same genetic material, but they don’t.
Method helps validate findings
In order to validate their findings, the researchers needed a way to tell whether an RNA was linear or circular since they share the same genetic code. This is where Batish’s expertise came in.
“You don’t ‘see’ RNA, per se, so you have to label it,” Batish said. “But, if you label it with something that is sequence-specific, it’s hard to tell if it’s linear or circular because the genetic code looks the same.”
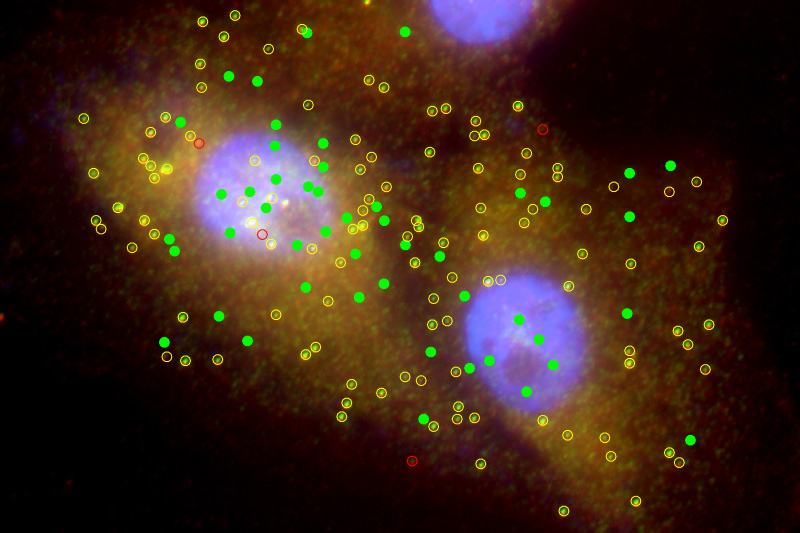
Batish had previously worked on probes that “light up” each RNA in an individual cell as a single bright spot under the fluorescence microscope to understand how biological systems operate on a cellular level. She adapted this method for distinguishing circular RNA from its linear RNA counterpart from the same gene by using a color combination method.
“Essentially, it’s like creating a pattern of beads on a necklace. Say the RNA we are working with contains red and green beads. We know that circular RNA is a closed circle of green beads only, so we add probes for both red and green beads and then image them under a fluorescence microscope,” said Batish. “If we see a signal for both red and green at the same spot, which appear as yellow (combination of green and red) in the sample, we know it is linear RNA. If it doesn’t have red, it must be circular RNA.”
This method allowed them to simultaneously visualize linear and circular RNA within a single cell.
“This is the first time that we have realized that RNA with the same genetic sequence can sometimes perform two roles, in this case both as a cancer suppressor and a cancer promoter, and that this change in role occurs at the RNA-level,” said Batish. “The identification of this new genetic unit opens up new opportunities to understand the genetics of cancer and the role of circRNA in cancer biology.”
And because a unique junction is created where the ends of circRNA come together, Batish said they may be able to develop treatment protocols to uniquely target the circular RNA, but leave the linear RNA alone. This could provide a way to target treatment to stop the circular RNA from turning off the cancer-suppressing effect in the body.
So, what’s next for Batish?
While this research study focused on connective and soft tissue tumors or diseases like mesenchymal tumors, Batish said the technique developed in her lab could be used on any cancer, because every cancer has circular RNA.
Batish plans to conduct experiments to see if what they have observed at the cellular level also occurs in tissue samples. Studying this expression in both healthy and diseased tissue, she said, would help her better understand the biosignature of circular RNA.
“If we can show that it persists in samples that are left out and not treated properly, that has real value. This is because circular RNA is differentially expressed, meaning my lungs will express different circular RNA than my brain and other tissues or organs,” said Batish.
“So imagine if you have that biosignature, and you can draw blood from a patient and look at what circular RNAs they have, you might be able to identify what kind of cancer a person has by these cancer markers instead of sending the patient for imaging or other testing. People are working on this research, so we’ll see.”
Batish also wants to study whether circular RNA found in tumors is present in cell signaling molecules known as extracellular vesicles. She describes these vesicles as letters, FedEx packages that cells put together and deliver to neighboring cells to tell them what’s going on nearby.
“One thought is that cancer may actually be hijacking this delivery system, adding ‘fake news’ into the packages that tell the neighboring cells that all is okay in their cell, while creating a microenvironment in which the cancer can grow,” she said.
Since every cancer starts with one single cell, Batish wants to explore the role circular RNA may play in this messaging. It might provide a pathway to understand how cell-to-cell communication is used by tumor cells.
She also wants to develop tools to enable live imaging of circular RNA in cells. Working in collaboration with Jeff Caplan, director of the Bio-Imaging Center at the Delaware Biotechnology Institute, Batish is exploring ways to add a “tracking device” of sorts into the cell that would allow them to follow the signal in real-time as circular RNA is formed.
“That would be really groundbreaking if we can do it,” said Batish.

Contact Us
Have a UDaily story idea?
Contact us at ocm@udel.edu
Members of the press
Contact us at 302-831-NEWS or visit the Media Relations website

